本文主要是介绍signature=70e902f89f03293bdcc59e78538792ab,Imaging EGFR and HER3 through,希望对大家解决编程问题提供一定的参考价值,需要的开发者们随着小编来一起学习吧!
Characterization of 89Zr-MEHD7945A
The labeling of MEHD7945A with 89Zr was straightforward. Radiolabeling yields of >95% were obtained with >99% purity after purification. A specific activity of 4.53 ± 0.65 mCi/mg (25.5 ± 3.7 MBq/nmol) was established. The labeled protein retained its immunoreactivity toward both EGFR and HER3 with 74 ± 0.5% (n = 3) retention, which is within range of acceptable immunoreactivities (>60%) for clinical use89Zr-MEHD7945A remains moderately intact >94% in both saline and 1:1 human serum:saline, over a 120 h incubation period at 37 °C (Supplementary Fig. S1).
EGFR and HER3 expression in established pancreatic cancer cells
Among the three pancreatic cell lines, AsPC-1 (Supplementary Fig. S2A) displayed the highest EGFR and HER3 staining with ~85% of the cell population co-expressing both receptors. BxPC-3 (Supplementary Fig. S2B) demonstrated approximately ~74% of the cell population staining for both receptors. A very low level of Mia PACA2 (Supplementary Fig. S2C) cells co-express both receptors (0.42%). Western blots demonstrated relatively equal expression of EGFR between AsPC-1 and BxPC-3 cell lines, with almost no EGFR expression in Mia PACA2 (Supplementary Fig. S2D). The HER3 order of expression for the three pancreatic cells are as follows: AsPC-1 > BxPC-3 > Mia PACA2.
In vitro internalization studies
Internalization of 89Zr-MEHD7945A in all cell lines was conducted at 37 °C (Fig. 1A, left). BxPC-3 displayed the highest uptake from 3.29 ± 0.28% at 1 h to almost a two-fold increase at 24 h with 5.91 ± 0.05% of the tracer internalized. In AsPC-1, the tracer was steadily internalized over time (2.58 ± 0.23% at 1 h, 3.23 ± 0.26% at 4 h and 4.70 ± 0.52% at 24 h) while the negative control cell line, Mia PACA2 demonstrated lower uptake across all time points (1.15 ± 0.06% at 1 h, 1.76 ± 0.17% at 4 h and 3.04 ± 0.21% at 24 h respectively). Significantly lower accumulation was observed in all cell lines at 4 °C, supporting an endocytotic mechanism of internalization (Fig. 1A, right).
Figure 1

In vitro characterization. 89Zr-MEHD7945A demonstrated higher internalization rates in BxPC-3 and AsPC-1 at 37 °C (left) compared to the EGFR/HER3-negative Mia PACA2 pancreatic cancer cell line. 89Zr-MEHD7945A showed a decrease in internalization at 4 °C (right) in all cell lines. (A) 89Zr-MEHD7945A demonstrated successful blocking with cold MEHD7945A and cetuximab at 10× and 25× doses. Blocking with DL3.6b at 10× lowered the uptake of the probe; binding was sustained at 25× dose of the anti-HER3 mAb. In both AsPC-1 and BxPC-3. (B) Non-linear regression analysis determined two sets of KD and Bmax for AsPC-1 with an. (C) The KD and Bmax values for BxPC-3 (D) are within the same range as the established values in AsPC-1 with an IC50 ~ 0.37 nM. (*Denote p
In vitro competitive inhibition studies
In AsPC-1 cells, (Fig. 1B and Supplementary Table S1) co-administration of 89Zr-MEHD7945A and unlabeled MEHD7945A at an excess of 10- and 25-fold for 1 h reduced uptake (1.97 ± 0.74%, p
In BxPC-3 cells, normal uptake of the radiotracer was measured at 11.72 ± 3.54% (Fig. 1B and Supplementary Table S1). Binding was significantly reduced when co-administered with MEHD7945A (10-fold: 1.02 ± 0.34%, p = 0.0001; 25-fold: 0.29 ± 0.16%, p = 0.0001) and cetuximab (10-fold: 1.21 ± 0.46%, p = 0.0001; 25-fold: 1.02 ± 0.23%, p = 0.0001). Competition with 10-fold DL3.6b lowered radiotracer accumulation to 6.98 ± 0.69% (p = 0.0001). Interestingly, radiotracer uptake was not diminished at 25-fold excess DL3.6b with 16.36 ± 0.88% binding (p = 0.0001). From the flow cytometry analysis, control cells exhibited 307.7 ± 11.2 MFI and 127.0 ± 3.6 MFI for EGFR and HER3 respectively. MEHD7945A-blocked cells decreased EGFR expression (182.7 ± 28.4 MFI, p = 0.0001) and HER3 (95.4 ± 7.7 MFI, p = 0.0123). Exposure to cetuximab mitigated EGFR expression (3.3 ± 0.8 MFI, p = 0.0001) whereas HER3 abundance remained unchanged (130.3 ± 4.0 MFI). Incubation with DL3.6b decreased HER3 (63.3 ± 8.2 MFI, p = 0.0001) while EGFR levels remained similar to unblocked control (284.0 ± 13.1 MFI, p = 0.0739) (Supplementary Fig. S3 and Supplementary Table S1).
In vitro binding affinity
Surface plasmon resonance (SPR) analysis of the dissociation constant (KD) of DFO-derivatized MEHD7945A vs. the unmodified mAb demonstrated relatively similar binding for HER3 (37 pM vs. 8.1 pM, respectively) and EGFR (3.4 nM vs. 1.9 nM respectively). Competitive binding of 89Zr-MEHD7945A with increasing concentrations of cold MEHD7945A was conducted in both BxPC-3 and AsPC-1 cells. In AsPC-1 (Fig. 1C, left), a KDHi ~ 0.31 nM and KDLo ~ 29.00 nM were achieved. Measured BmaxHi and BmaxLo values were ~1.94 × 105 and ~7.02 × 104 receptors sites were available. The IC50 was ~0.37 nM (Fig. 1C, right). From the plot in Fig. 1D (left), the dissociation constants KDHi and KDLo in BxPC-3 cells were determined to be 0.34 nM and 12.02 nM with a BmaxHi of ~7.25 × 104 and BmaxLo of ~1.10 × 105. The IC50 for 89Zr-MEHD7945A was observed to be ~0.48 nM in BxPC-3 (Fig. 1D, right).
In vivo EGFR/HER3-PET Imaging
In Fig. 2A, cumulative tumor uptake of 89Zr-MEHD7945A was observed in the AsPC-1 xenograft with 3.98 ± 0.21 percent injected dose per gram of tissue (%ID/g) at 24 h post-injection (p.i.) with the maximum tumor-bound activity achieved at 72 h p.i. at 6.18 ± 1.0%ID/g. Retention of the radiotracer was observed to as far as 96 h p.i. (6.23 ± 1.35%ID/g). AsPC-1 tumors imaged with the non-specific isotype 89Zr-IgG had significantly less uptake (0.62 ± 0.53%ID/g at 24 h, 0.80 ± 0.44%ID/g at 48 h, 1.03 ± 0.55%ID/g at 72 h, and 0.83 ± 0.29%ID/g, p
Figure 2

In vivo PET imaging. In AsPC-1 xenografts, tumor volumes-of-interest (VOI) expressed as % ID/g generated from the 89Zr-MEHD7945A PET scans exhibited uptake as early as 24 h p.i., peaking at 72 h p.i. and retained to as long as 96 h p.i. (A) 89Zr-IgG control PET scans showed minimal uptake within the tumor at all time points. (B) Similarly in BxPC-3 tumors, PET scans exhibited uptake at 24 h p.i. with a peak at 72 h p.i. (C) Non-specific tumor uptake using 89Zr-IgG in BxPC-3 xenografts showed nominal accumulation across all time points. (D) Whole body tissue distribution revealed high tumor tissue uptake of the tracer at 24 h p.i,, which plateaued at 48 h through 120 h p.i. in BxPC-3 xenografts. A competitive blocking study using unmodified MEHD7945A at 48 h p.i. displayed at least a two-fold decrease in tumor binding, indicative of the probe’s specificity. (E) Of note, normal pancreas demonstrated minimal non-specific binding on all time points, suggesting that an excellent signal-to-noise contrast can be achieved.
Whole Body Distribution of 89Zr-MEHD7945A
Tissue distribution of 89Zr-MEHD7945A was analyzed in BxPC-3 tumor-bearing mice at 24–120 h p.i. (Fig. 2E and Supplementary Table S2). Tumor uptake was detected at 24 h p.i. with 24.6 ± 6.7%ID/g. The accumulation reached 31.1 ± 3.9%ID/g at 48 h p.i. and plateaued over the later time points with 32.8 ± 4.7%ID/g at 96 h p.i. and 32.7 ± 5.3%ID.g at 120 h p.i. Tumor accumulation decreased by two-fold (13.0 ± 3.4%ID/g) at 48 h p.i when 89Zr-MEHD7845A was administered with an excess of unmodified antibody to compete and block the radiotracer for receptor sites. Uptake in normal pancreas was significantly lower with <2%ID/g across all time points. Other organs within close proximity to the pancreas demonstrated low non-specific accumulation. For example at 48 h p.i., non-targeted binding in the liver, the primary clearance route for most mAbs, was 2.5-fold less than tumor uptake at 9.7 ± 2%ID/g. At the same time points, tissue-uptake values for the spleen, stomach and gut were 3.7 ± 1.8%ID/g, 1.3 ± 0.7%ID/g and 1.2 ± 0.3%ID/g respectively. Bone accumulation was observed over time with 6.84 ± 3.85%ID/g at 48 h p.i., which plateaued at 8.86 ± 4.34%ID/g at 120 h p.i. Blocking with MEHD7945A at 48 h p.i. moderately lowered the bone uptake to 4.03 ± 1.60%ID/g but was statistically insignificant.
Ex Vivo Autoradiography and immunohistochemistry (IHC)
To understand the differences in tracer uptake between tumor models, spatial distribution of 89Zr-MEHD7945A was evaluated using digital autoradiography on AsPC-1 (Fig. 3A, left) and BxPC-3 tumors (Fig. 3A, right). Adjacent tumor sections of both AsPC-1 and BxPC-3 were evaluated for EGFR (Fig. 3B) and HER3 expression (Fig. 3C) by immunohistochemistry. Tissue viability and collagen content were qualitatively assessed via H&E staining (Supplementary Fig. S4A,B) and trichrome staining (Supplementary Fig. S4C), respectively. In both tumors, 89Zr-MEHD7945A accumulated in viable tumor regions with high EGFR expression (Fig. 3B). Staining of EGFR and HER3 showed strongest positivity on viable tumor cells. Although, the expression of EGFR in BxPC-3 and AsPC-1 tumor sections was similar, there was higher expression of HER3 in AsPC-1 when compared to BxPC-3. Areas with strong staining for HER3 co-localized with areas of high 89Zr-MEHD7945A registration (Fig. 3C). Taken together, the localization pattern indicates the dependence of 89Zr-MEHD7945A on both target expression and local pharmacokinetics. Collagen expression was qualitatively equivalent for both tumor sections. Looking at cell density, BxPC-3 tumors were observed to be two-fold more “cell dense” than AsPC-1 xenografts (258.8 ± 64.2 cells vs. 71.3 ± 14.9 cells, p = 0.0013, (Supplementary Fig. S4D).
Figure 3

Autoradiography and histology. Autoradiographs (A) of excised AsPC-1 (left) and BxPC-3 (right) tumor sections depicted co-localization of the tracer in areas where EGFR (BxPC-3 3+, AsPC-1 2+) (B) and HER3 (BxPC-3 1+, AsPC-1 2+) (C) are expressed.
Assessment of EGFR and/or HER3 dual specificity of 89Zr-MEHD7945A
To determine whether 89Zr-MEHD7945A binds to either EGFR or HER3 alone or both, in vivo blocking studies were conducted. Blocking EGFR with cetuximab in AsPC-1 tumors (Fig. 4A and Supplementary Table S3) lowered the uptake of 89Zr-MEHD7945A by almost 3-fold (19.30 ± 5.60%ID/g vs. 7.72 ± 6.70%ID/g, p = 0.0457). In contrast, blocking the HER3 epitope with DL3.6b did not alter tumor accumulation of 89Zr-MEHD7945A (18.79 ± 8.75%ID/g).
Figure 4

In vivo competitive inhibition. In AsPC-1 xenografts, blocking with cetuximab (EGFR block) showed an almost 2-fold decrease in 89Zr-MEHD7945A uptake, whereas blocking HER3 with DL3.6b did not change probe uptake. (A) In BxPC-3 xenografts, blocking with cetuximab (EGFR block) showed a slight decrease in 89Zr-MEHD7945A, whereas blocking with DL3.6b (HER3 block) showed a statistically significant, increase in probe accumulation. (B) IHC staining in BxPC-3 tumors blocked with 25× cetuximab (25x EGFR, left), 25x DL3.6b (25x HER3, middle) or left unblocked (right) were assessed by IHC for EGFR (top) and HER3 (bottom) expression, and showed an increase in EGFR and HER3 in both blocked cohorts. (C) Tumor sections depicted for IHC are shown in 100×. Densitometry analysis of western blots on tumor lysates (n = 2) from AsPC-1 (left) and BxPC-3 (right) that were untreated, exposed to EGFR-block with cetuximab and a HER3-block with DL3.6b. (D) Densitometry is shown as a ratio of target protein/loading control.
In BxPC-3 xenografts (Fig. 4B and Supplementary Table S3), exposure to cetuximab showed no difference in 89Zr-MEHD7945A uptake between control (28.56 ± 10.81%ID/g) and cetuximab-blocked tumors (21.79 ± 14.19%ID/g, p = 0.39). Competition of the tracer with DL3.6b demonstrated an approximately two-fold increase in radiotracer accumulation of 46.89 ± 20.1%ID/g (p = 0.0021).
We next determined the underlying mechanism for the unexpected trend in tumor uptake of the blocked cohorts. Pathological analysis revealed an increase in EGFR (Fig. 4C, top) and HER3 (Fig. 4C, bottom) staining in both EGFR and HER3 blocked BxPC-3 tumors compared to control. From the western blots, cetuximab-treated AsPC-1 exhibited a two-fold decrease in total EGFR protein, coupled with a 1.3-fold increase in HER3 as shown by densitometry (Fig. 4D). With the DL3.6b blocked AsPC-1 tumors, a 2.25-fold increase in EGFR with a concomitant three-fold decrease in HER3 was displayed. Similar trends were observed in BxPC-3 wherein cetuximab-blocked groups displayed suppressed EGFR and HER3 (Fig. 4D). Saturation of the HER3 sites exhibited a moderate increase in EGFR expression, coupled with a decrease in HER3. These results imply that the blocking dose of antibody (cetuximab or DL3.6b) may be inducing an upregulation of either receptor in an attempt to maintain proliferation.
Ex vivo competitive binding assay
Figure 5A showed that an addition of excess cold MEHD7945A (100 nM) blocks the binding of 89Zr-MEHD7945A (1 nM). This demonstrated a saturable, concentration-dependent binding of the tracer. From the Scatchard plot (Fig. 5B), the Bmax was found to be 336.0 ± 25.5 fmol/mg or 2 × 105 binding sites per cell (assuming 1 million cells per tumor) and the KD was found to be ~0.51 ± 0.12 nM.
Figure 5
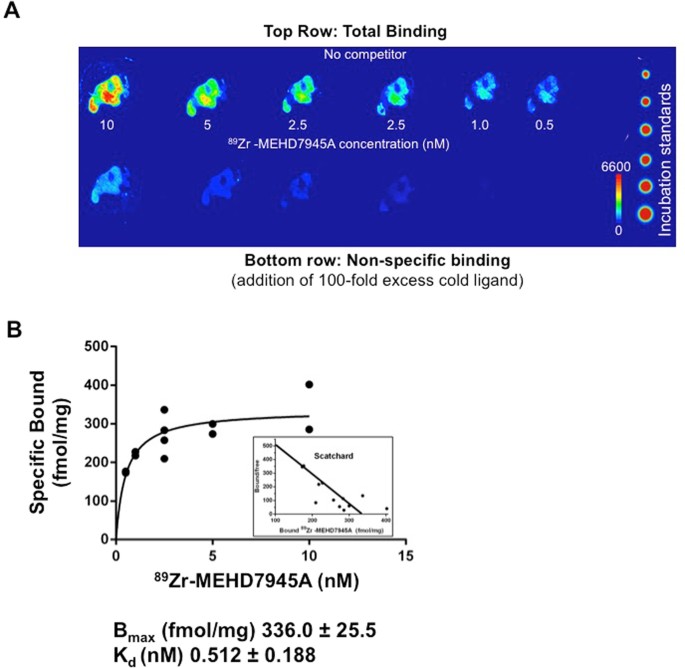
Ex vivo competitive binding assay. A digital autoradiograph of AsPC-1 tumor sections displayed saturable, concentration-dependent binding of 89Zr-MEHD7945A upon addition of 100-fold excess cold MEHD7945A. (A) A non-linear regression analysis and Scatchard plot of 89Zr-MEHD7945A plotted against the amount of bound ligand shows the Bmax ~ 336 fmol/mg and a KD ~ 0.51 nM (B).
这篇关于signature=70e902f89f03293bdcc59e78538792ab,Imaging EGFR and HER3 through的文章就介绍到这儿,希望我们推荐的文章对编程师们有所帮助!